slideWindowTunePlot - slideWindowTunePlot - plotSlideWindowTune backward compatability
Description¶
Wrapper function for plotSlideWindowTune
Usage¶
slideWindowTunePlot(
tuneList,
plotFiltered = c(TRUE, FALSE, NULL, "filtered", "remaining", "per_mutation"),
percentage = FALSE,
jitter.x = FALSE,
jitter.x.amt = 0.1,
jitter.y = FALSE,
jitter.y.amt = 0.1,
pchs = 1,
ltys = 2,
cols = 1,
plotLegend = TRUE,
legendPos = "topright",
legendHoriz = FALSE,
legendCex = 1,
title = NULL,
returnRaw = FALSE
)
Arguments¶
- tuneList
- a list of logical matrices returned by slideWindowTune.
- plotFiltered
- whether to plot the number of filtered (
TRUEorfiltered), or remaining (FALSE or remaining) sequences for each mutation threshold. UseNULLorper_mutationto plot the number of sequences at each mutation value. Default isTRUE. - percentage
- whether to plot on the y-axis the percentage of filtered sequences
(as opposed to the absolute number). Default is
FALSE. - jitter.x
- whether to jitter x-axis values. Default is
FALSE. - jitter.x.amt
- amount of jittering to be applied on x-axis values if
jitter.x=TRUE. Default is 0.1. - jitter.y
- whether to jitter y-axis values. Default is
FALSE. - jitter.y.amt
- amount of jittering to be applied on y-axis values if
jitter.y=TRUE. Default is 0.1. - pchs
- point types to pass on to plot.
- ltys
- line types to pass on to plot.
- cols
- colors to pass on to plot.
- plotLegend
- whether to plot legend. Default is
TRUE. - legendPos
- position of legend to pass on to legend. Can be either a
numeric vector specifying x-y coordinates, or one of
"topright","center", etc. Default is"topright". - legendHoriz
- whether to make legend horizontal. Default is
FALSE. - legendCex
- numeric values by which legend should be magnified relative to 1.
- title
- plot main title. Default is NULL (no title)
- returnRaw
- Return a data.frame with sequence counts (TRUE) or a
plot. Default is
FALSE.
Details¶
For each windowSize, if plotFiltered=TRUE, the x-axis
represents a mutation threshold range, and the y-axis the number of
sequences that have at least that number of mutations. If
plotFiltered=TRUE, the y-axis represents the number of sequences
that have less mutations than the mutation threshold range. For the same
window size, a sequence can be included in the counts for different
mutation thresholds. For example, sequence “CCACCAAAA” with germline
“AAAAAAAAA” has 4 mutations. This sequence has at least 2 mutations
and at least 3 mutations, in a window of size 4. the sequence will
be included in the sequence count for mutation thresholds 2 and 3.
If plotFiltered=TRUE, the sequences are counted only once for
each window size, at their largest mutation threshold. The above
example sequence would be included in the sequence count for
mutation threshold 3.
When plotting, a user-defined amount of jittering can be applied on values plotted
on either axis or both axes via adjusting jitter.x, jitter.y,
jitter.x.amt and jitter.y.amt. This may be help with visually distinguishing
lines for different window sizes in case they are very close or identical to each other.
If plotting percentages (percentage=TRUE) and using jittering on the y-axis values
(jitter.y=TRUE), it is strongly recommended that jitter.y.amt be set very
small (e.g. 0.01).
NA for a combination of mutThresh and windowSize where
mutThresh is greater than windowSize will not be plotted.
Examples¶
# Use an entry in the example data for input and germline sequence
data(ExampleDb, package="alakazam")
# Try out thresholds of 2-4 mutations in window sizes of 3-5 nucleotides
# on a subset of ExampleDb
tuneList <- slideWindowTune(db = ExampleDb[1:10, ],
mutThreshRange = 2:4, windowSizeRange = 3:5,
verbose = FALSE)
|
| | 0%
|
|======= | 10%
|
|============== | 20%
|
|===================== | 30%
|
|============================ | 40%
|
|=================================== | 50%
|
|========================================== | 60%
|
|================================================= | 70%
|
|======================================================== | 80%
|
|=============================================================== | 90%
|
|======================================================================| 100%
|
| | 0%
|
|======== | 11%
|
|================ | 22%
|
|======================= | 33%
|
|=============================== | 44%
|
|======================================= | 56%
|
|=============================================== | 67%
|
|====================================================== | 78%
|
|============================================================== | 89%
|
|======================================================================| 100%
# Visualize
# Plot numbers of sequences filtered without jittering y-axis values
slideWindowTunePlot(tuneList, pchs=1:3, ltys=1:3, cols=1:3,
plotFiltered=TRUE, jitter.y=FALSE)
Warning:slideWindowTunePlot() is deprecated, please see plotSlideWindowTune() for future use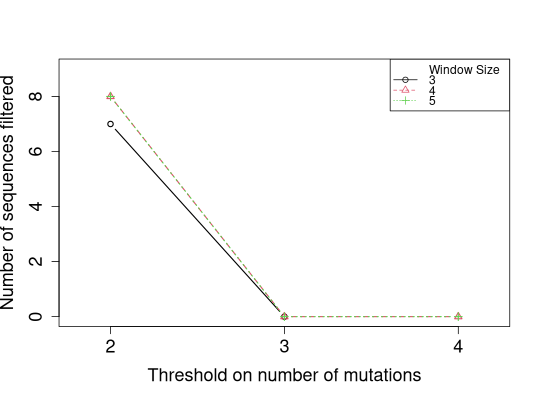
# Notice that some of the lines overlap
# Jittering could help
slideWindowTunePlot(tuneList, pchs=1:3, ltys=1:3, cols=1:3,
plotFiltered=TRUE, jitter.y=TRUE)
Warning:slideWindowTunePlot() is deprecated, please see plotSlideWindowTune() for future use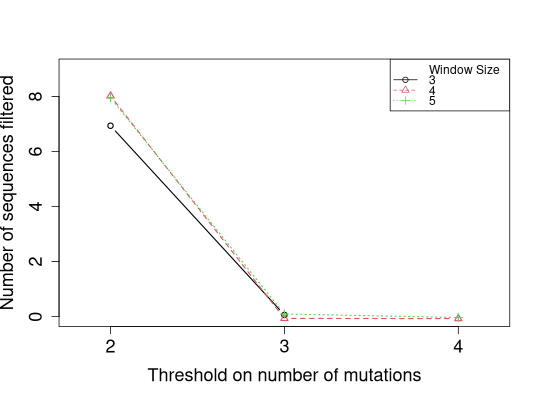
# Plot numbers of sequences remaining instead of filtered
slideWindowTunePlot(tuneList, pchs=1:3, ltys=1:3, cols=1:3,
plotFiltered=FALSE, jitter.y=TRUE,
legendPos="bottomright")
Warning:slideWindowTunePlot() is deprecated, please see plotSlideWindowTune() for future use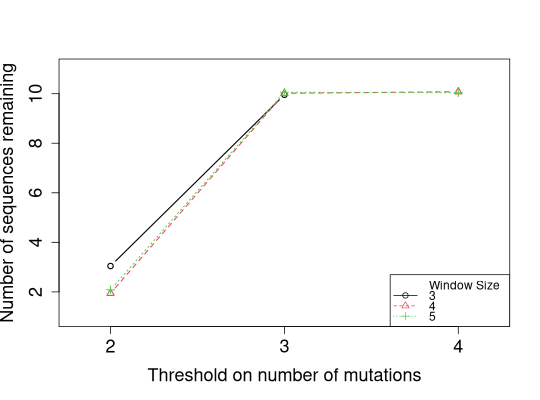
# Plot percentages of sequences filtered with a tiny amount of jittering
slideWindowTunePlot(tuneList, pchs=1:3, ltys=1:3, cols=1:3,
plotFiltered=TRUE, percentage=TRUE,
jitter.y=TRUE, jitter.y.amt=0.01)
Warning:slideWindowTunePlot() is deprecated, please see plotSlideWindowTune() for future use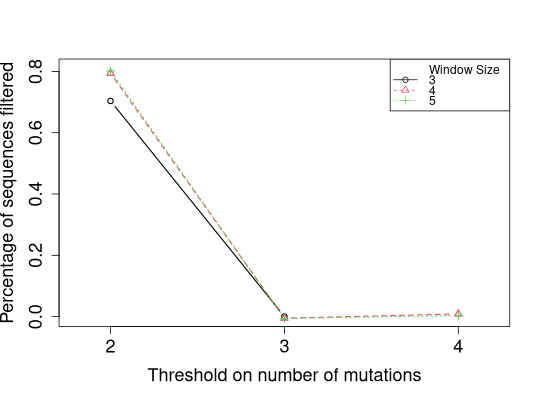
See also¶
See slideWindowTune for how to get tuneList. See jitter for
use of amount of jittering.